This post is a tribute to the work of my students in the subject “Basic Python Programming for Scientists“, as they did a great job in their final projects I want to show you here some examples that could be useful for many Python beginners. Remember that you can download the course slides from Didactic Materials sections.
First of all, I want so say that it was my first experience as Python Programming teacher and I was gratefully surprised with my students. I have learned a lot with them and I can say that some of them outperformed the teacher. I also want to thank to Medhat Helmy and Piotr Bentkowski for helping me in the teaching task. Unfortunately I cannot have the same opinion from the Adam Mickiewicz University that didn’t pay me any money for teaching, welcome to Poland…
Today I’ll show you 2 projects that overcame all my expectations: Image Quiz from Błażej Szymański and Temperature converter by Michal Stachowiak.. In next days I’ll publish more incredible projects.
Image Quiz
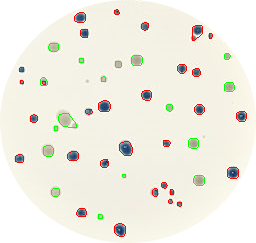
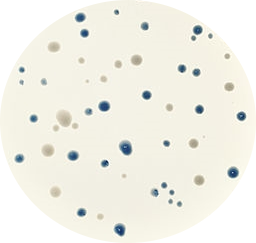
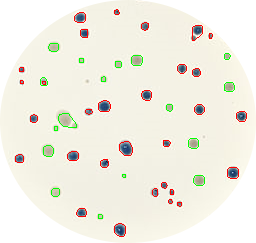
Maybe the project that impressed me the most was ‘Image Quiz’ by Błażej Szymański. He was planning at the beginning to write a python script that automatically downloads different airplane model images from Google, and later use these images with an HTML+JavaScript interface to play a quiz to guess which is the correct airplane model shown in a randomly selected image. I suggested some minor changes, like using a more biological topic as bird species instead of airplanes. And now you can check yourself the final result playing the Image Quiz online!
The project consists basically in 4 python scripts:
- fetcher.py: reads a file ‘list.txt’ containing the names of the birds to look for and use the Google API to retrieve the bird photo URLs that are stored as JSON format in ‘data.txt’.
- saver.py: retrieves the image files and saves them at ‘pictures’ folder using their URLs stored in ‘data.txt’.
- resizer.py: checks if the images are larger than 800px, if so they are resized.
- final.py: saves a dictionary with bird names as keys and arrays with the image locations as values in JSON format into ‘result.txt’. This JSON file will be used later by a JavaScript function to prepare the quiz.
fetcher.py:
import requests
import json
import os
result = []
question = []
with open("list.txt","r") as file:
for line in file:
question.append(line[:-1])
for q in question:
stuff = {"key":"WRITE HERE YOUR GOOGLE API KEY",
"num":10,
"searchType":"image",
"q":q,
"cx":"003535310068073691339:ebzdlw5y6gq",
"excludeTerms":"stock" #to get rid of stock images
}
j = requests.get("https://www.googleapis.com/customsearch/v1", params=stuff)
r = json.loads(j.text)
x = []
for each in r["items"]:
x.append(each["link"])
result.append(x)
dictionary = dict(zip(question,result))
with open('data.txt', 'w') as resultfile:
json.dump(dictionary, resultfile)
saver.py:
import os
import requests
import shutil
import json
with open("data.txt","r") as data:
dictionary = json.load(data)
for key in dictionary.keys():
for index, each in enumerate(dictionary[key]):
r = requests.get(each, stream=True)
folder = "./pictures/{pic}".format(pic=key)
os.makedirs(folder, exist_ok=True)
print("Retrieving image {} from {}".format(index+1,key))
with open(folder + "/" + str(index) + ".jpeg", "wb") as file:
shutil.copyfileobj(r.raw, file)
print("\nPlease review downloaded pictures and delete invalid ones.")
resizer.py:
import os
from PIL import Image
for thing in os.listdir("./pictures"):
for model in os.listdir("./pictures/" + thing):
if model == "Thumbs.db": #to avoid windows thumbnails
continue
im = Image.open("./pictures/" + thing + "/" + model)
if im.size[1] > 800:
ratio = im.size[0]/im.size[1]
x = ((round(ratio*800)),800)
print(model,"resized to",x)
im = im.resize(x,Image.ANTIALIAS)
im.save("./pictures/" + thing + "/" + model)
final.py:
import os
import json
keys = []
temp = []
values = []
for thing in os.listdir("./pictures"):
keys.append(thing)
for file in os.listdir("./pictures"+"/"+thing):
if file != "Thumbs.db":
temp.append("./pictures"+"/"+thing+"/"+file)
values.append(temp)
temp=[]
dictionary = dict(zip(keys,values))
with open("result.txt", "w") as result:
json.dump(dictionary, result)
Temperature converter
‘Temperature converter’ by Michal Stachowiak, is a script that converts temperature values in different scales and saves the results in memory.
He implemented temperature conversions between Kelvin, Celsius and Fahrenheit scales together with a database to store and modify the results. Here I’ll show a reduced version of the script that only converts from Celsius to Kelvin.
#################################
#FUNCTIONS FOR CONVERTING
def C_on_K (C):
K = C + 273.15
return K
########################################
#MAIN PROGRAM
print("\n\n")
print(" ===========================")
print(" Temperature Converter")
print(" ===========================\n\n")
print("Input and output are stored in a database\n\n")
gearbox = 0
index = 0
while gearbox < 100: #You can execute this program 100 times print("Choose the converter:") print("----------------------") print(" 1. C => K")
print("----------------------")
print("Chose the action:")
print("----------------------")
print(" 7. Show all records")
print(" 8. Show specific record")
print(" 9. Edit specific record")
print("10. Delete all records")
print("11. Delete specific record")
print("12. Close the program")
print("----------------------")
print("\nChoice 1-12:")
answer = input()
str(answer)
######################################################
if answer == "1":
controler = 1
while controler == 1:
print ("Provide the temperature in Celsius degree:")
C = float(input())
if C < -273: #there is no C temp below that value, so ask again for proper temperature
print ("There is no temperature in C degree below -273")
print ("Please provide proper C temperature")
controler = 1
else:
equation5 = C_on_K(C)
print("=========================")
print( "{:.2f}C degree is {:.2f}K".format(C,equation5))
print("=========================")
controler = 0
gearbox += 1
print("Press any key to continue...")
input()
elif answer == "12":
print("Thank You for using Temperature Converter")
print("If You enjoy the program, I appreciate any donations")
print("Press any key to quit...")
input()
sys.exit(0)
else:
print("Please provide number from 1 to 12")
gearbox += 1


 The didactic materials in this new section will be licensed as
The didactic materials in this new section will be licensed as 

